USE CASES
Explore diverse use cases of the automated SISCAPA workflow, from oncology and infectious disease diagnostics to neurological and cardiovascular health monitoring, utilizing targeted peptide enrichment to enhance sensitivity, specificity, and multiplexing capabilities for precise protein measurement.
CLICK ON THE USE CASE TO THE RIGHT THAT BEST DIRECTS YOUR INTEREST.
ONCOLOGY
Accurate quantitation of protein targets (e.g., PD-1, BTK), drugs (e.g., anti-PD-1 monoclonals, covalent kinase inhibitors), and biomarkers (e.g., MEK phosphorylation) is critical in many areas of oncology.
PROTEIN DRUGS, FREE VS. BOUND FORMS
Many protein drugs contain unique proteolytic peptide sequences – following sample digestion, these allow accurate measurement of total drug without interferences caused by binding of receptors, antibodies, etc. to the drug.
In many cases, peptides can be identified in the complementarity determining regions (CDRs) of therapeutic monoclonal antibodies that are unique to the drug and absent from any other endogenous protein. Measurement of these peptides by SISCAPA-MS ensures an interference-free measurement of total drug.
In addition to measuring total amounts of proteins, we have developed workflows to measure free-vs-bound forms of targets and drugs, as well as complexes of drugs with endogenous autoantibodies.
Clinical samples can be fractionated prior to total drug measurement into free vs complexed sub-samples. For example, free soluble PD-L1 (sPD-L1) can be can be captured and removed from a sample by magnetic beads coated with Avelumab (an anti-PD-L1 antibody), generating two sub-samples. Measurement of both sPD-L1 and Avelumab in both samples allows calculation of total drug (Avelumab), total sPD-L1, and the amount of Avelumab complexed with sPD-L1 (the “bound” drug fraction).
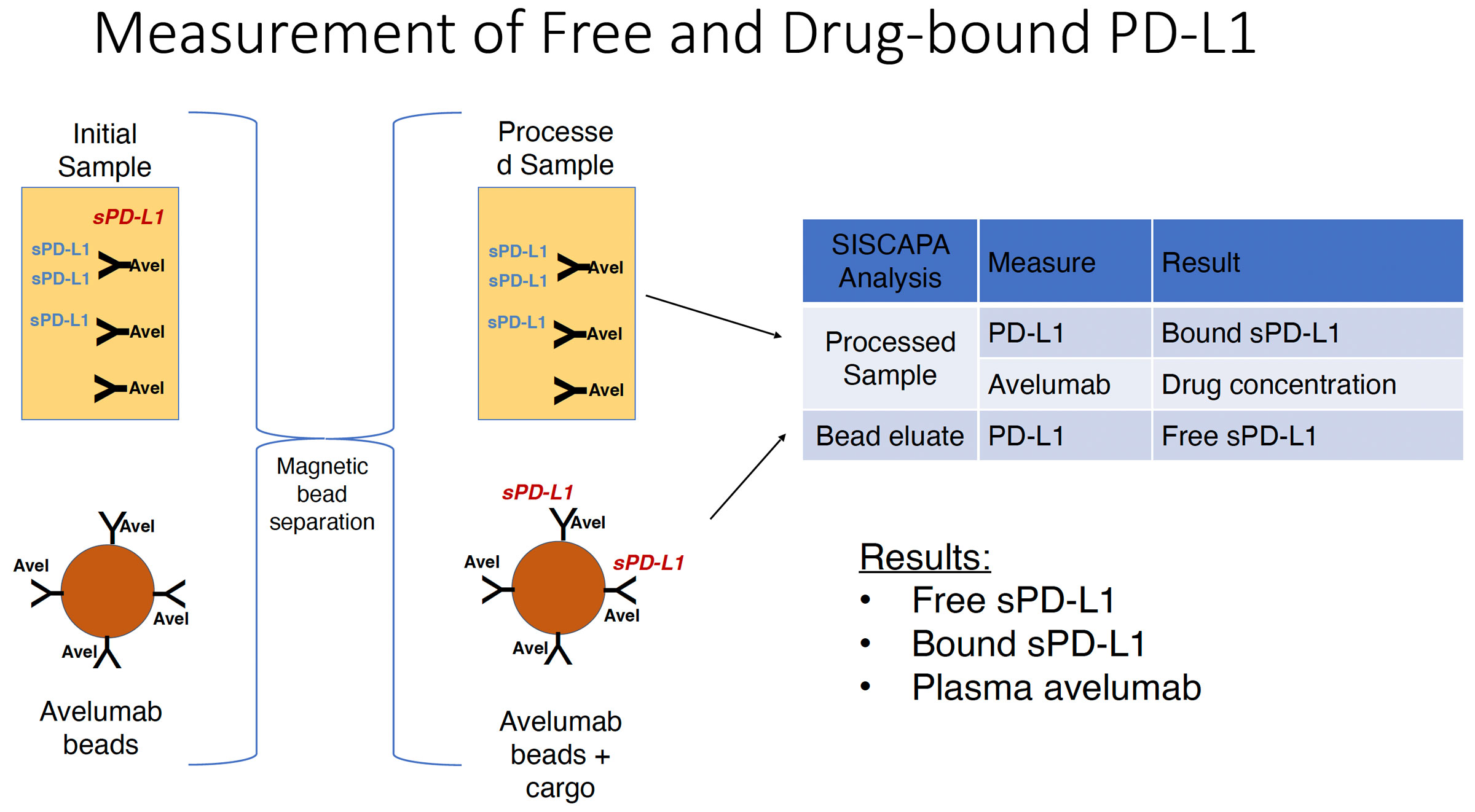
Similarly, samples can be fractionated to separate molecules of an antibody drug whose active sites are blocked by interaction with an endogenous anti-drug antibody (ADA), allowing determination of the total amount of drug and the fraction that is active.
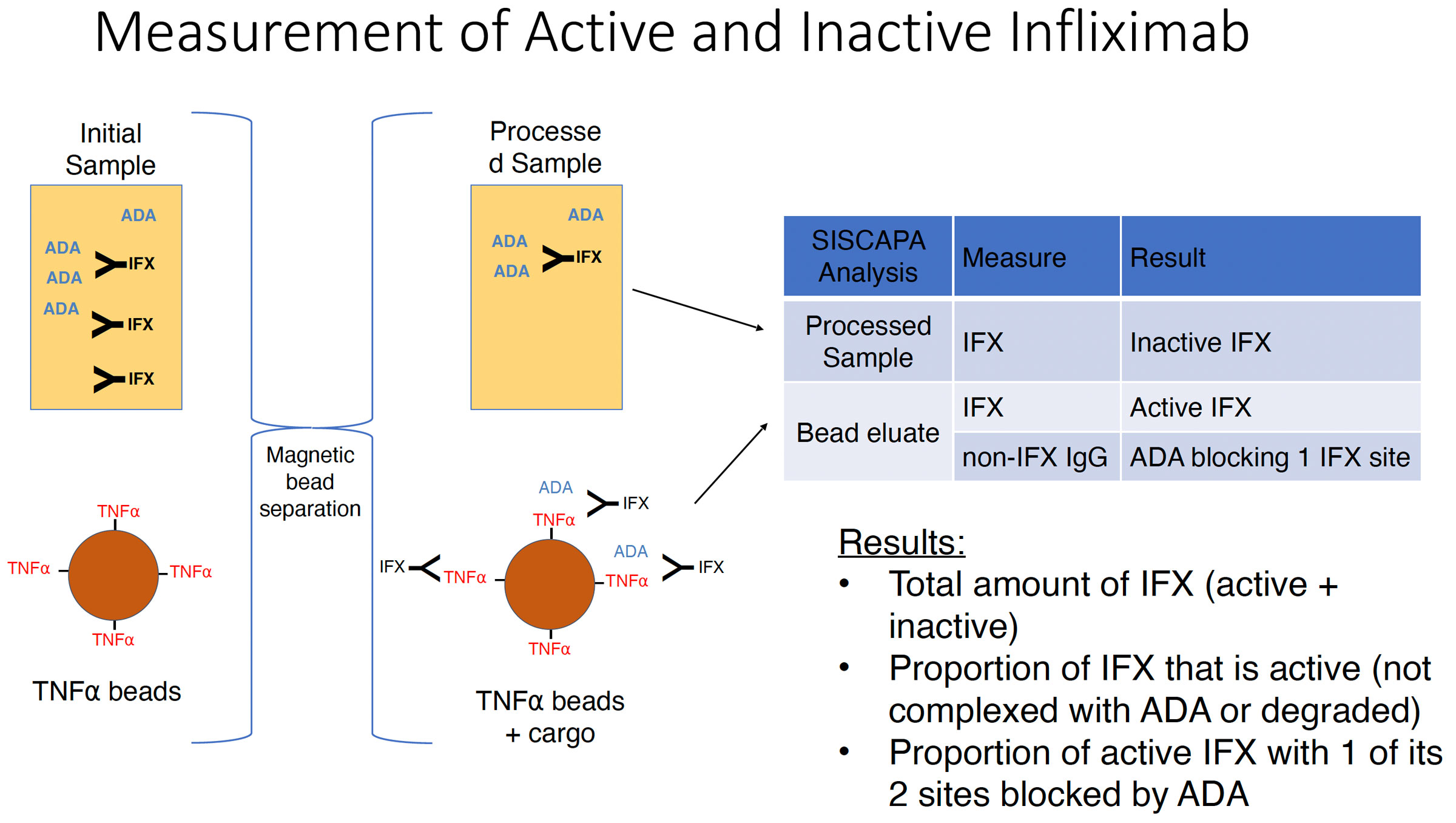
Contact us about your specific free-vs-bound challenge.
DIAGNOSTIC AND PROGNOSTIC BIOMARKERS
Several prognostic protein panels widely used in oncology practice (the Glasgow Prognostic Score [GPS] and the neutrophil:lymphocyte ratio [LNR]) are composed of individual markers included in our general inflammation panel. Application of these panels in longitudinal home-collected dried blood spots presents an important addition to patient monitoring in oncology clinical trials. In addition, assays for most of the FDA-cleared cancer diagnostic markers (e.g., AFP, CEA, PSA, mesothelin, etc.) are included in our catalog menu and oncology panel.
INFECTIOUS DISEASE
Infectious organisms produce proteins not normally present in a host, and detection of these at any level indicates a likely infection.
Based on experimental evidence (e.g., from proteomics) or in silico predictions (from organism genomes), organism-specific peptides can be selected as SISCAPA targets, and used for definitive diagnosis of infection.
In this application, the benefits of using SISCAPA include:
- A large increase in sensitivity (arising from enrichment of target peptides and depletion of matrix)
- Absolute specificity of MRM-MS detection (providing a significant improvement over lateral flow point of care antigen tests)
- Resistance to viral mutations (a panel of peptide targets drastically reduces the chance that any one mutation can render the assay blind)
During the COVID pandemic, we undertook development of a comprehensive suite of definitive SISCAPA assays for the SARS-CoV-2 virus. The project focused initially on 10 tryptic peptides from the viral NCAP protein (NCAP was chosen because it is present in many copies per virion and it demonstrates low rates of mutation compared to the Spike protein). Monoclonal SISCAPA antibodies were then developed to 7 of the 10 initial targets, of which smaller panels of 2 or 3 assays were validated in clinical samples.
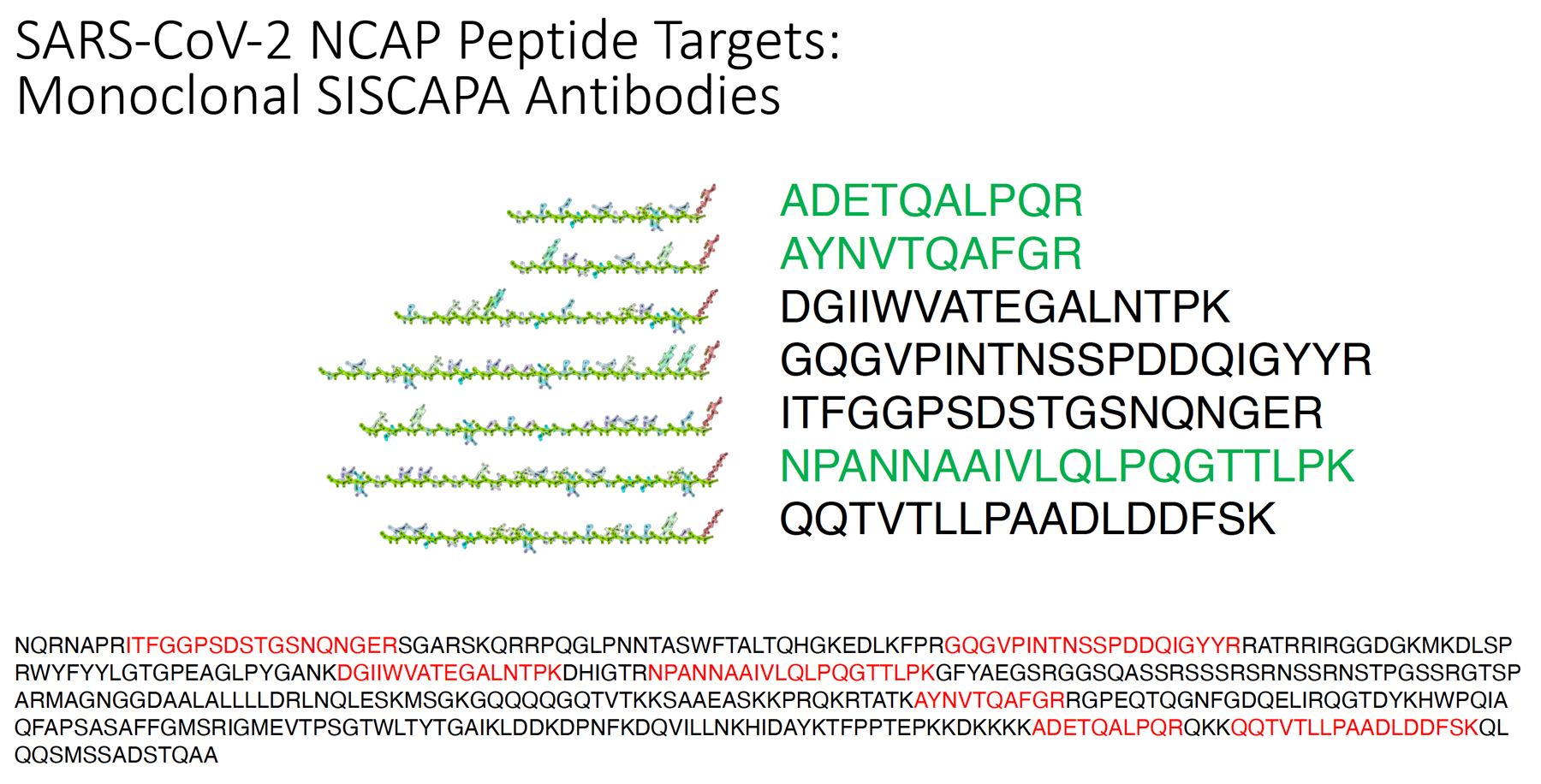

The COVID assays have been described in published studies with a range of collaborators and resulted in production of a commercial detection kit.
Additional projects currently underway include SISCAPA assays for proteins of syphilis, influenza A & B, and RSV.
Contact us to access SISCAPA COVID reagent
Operation Moonshot: rapid translation of a SARS-CoV-2 targeted peptide immunoaffinity liquid chromatography-tandem mass spectrometry test from research into routine clinical use. (2022) Clin Chem Lab Med 2023; 61(2): 302–310, https://doi.org/10.1515/cclm-2022-1000
GENE THERAPY
Genetic errors causing some heritable, often rare, diseases can now be corrected by introducing genes that restore in vivo expression of a functional version of a patient’s defective protein.
Clearly demonstrating expression of a repaired gene product is a key component of the evaluation of gene therapies. In most cases, it is useful to accurately measure and compare the levels of the repaired (functional) protein with that of the endogenous (defective) protein, with the goal of achieving a functionally significant replacement. Additionally, given the high degree of sequence homology between humans and animals used for pre-clinical studies, sequence-based specificity beyond that achievable with immunoassays and LBA’s is critical.
Given that the SISCAPA technology can develop assays against proteotypic peptide targets at single amino acid resolution, it has been successfully utilized for such repaired/endogenous protein comparisons, and cross-species assay development programs. Some of these assays are currently being used by clients for clinical and preclinical studies.
A similar example is described in the following publication.
INFLAMMATION
Inflammation is central to the understanding of human health and disease: while non-specific in a diagnostic sense, inflammation markers provide powerful tools for tracking disease and treatment responses over time.
CRP, the most frequently measured clinical inflammation indicator, is associated with negative developments in an astonishing range of health situations, including infection, arthritis, surgery, intense exercise, sleep apnea, depression, air pollution, welding fume exposure, Parkinson’s disease, pregnancy, inflammatory bowel disease, and various cancers, to name a few examples. While inflammation is thus clearly not disease-specific, changes in inflammation levels can serve as useful indicators of changing severity and efficacy of treatment in many disease areas. Many therapeutics that reduce inflammation are in use or under development. These usually act either directly (by targeting inflammation pathway signaling) or indirectly (by treating causes of inflammation).
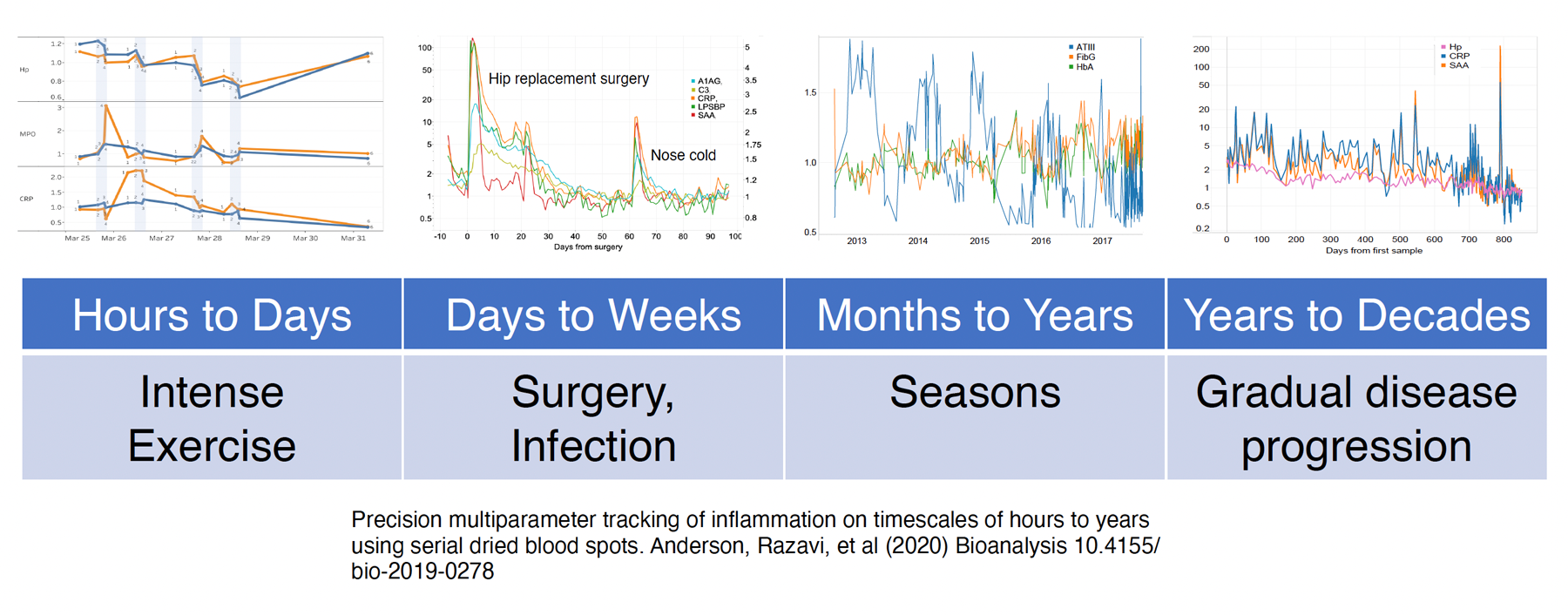
We have developed a broad panel of inflammation markers designed to detect responses on a range of timescales from hours to weeks – i.e., on clinically relevant timescales beyond the short-duration fluctuations of cytokines. These include major components of the acute phase response (CRP, SAA, orosomucoid, LPS-binding protein, mannose-binding lectin, haptoglobin, complement C3, and fibrinogen), IgM (indicator of an antibody immune response), and myeloperoxidase (indicator of neutrophil counts in whole blood). The panel is of particular value in following longitudinal changes within-patient – for example using finger-prick dried blood spot samples collection on a daily (or less-frequent) basis.
Precision multiparameter tracking of inflammation on timescales of hours to years using serial dried blood spots. L. Anderson et al., Bioanalysis, vol. 12, no. 13, pp. 937–955, Jul. 2020, doi: 10.4155/bio-2019-0278
NEUROLOGICAL DISEASE
Recent progress towards treatments for Alzheimers and Parkinson’s diseases, stroke and traumatic brain injury have highlighted the need for accurate measurement of protein indicators of neurological damage.
Neuronal proteins (e.g., Alpha-synuclein) released into body fluids represent useful diagnostic and surrogate biomarkers in neurological disease, albeit challenging to measure accurately due to their very low concentration. SISCAPA assays provide the interference-free enrichment required to detect low abundance molecules in dilute samples such as CSF, as well as complex matrices such as whole blood.
Neurological target proteins, such as membrane-embedded sodium channels involved in pain perception (e.g., Nav1.7 and Nav1.8) present specific measurement challenges because of their insolubility and low concentration. SISCAPA assays provide a means to enrich and detect these difficult targets using specific proteotypic peptides. [Figure on Nav1.7 and Nav1.8 – or better use the animation]
CARDIOVASCULAR DISEASE
Numerous protein biomarkers are used clinically as indicators of cardiovascular disease (CVD) risk, and these represent a natural opportunity to leverage panels of markers to achieve improved predictive value.
SISCAPA assays for the most basic CVD risk marker proteins (ApoA-I and ApoB lipoproteins) have been clinically validated as superior replacements for conventional HDL and LDL cholesterol measurements. A suite of additional SISCAPA assays for known CVD biomarkers (CRP, myeloperoxidase, Lp(a), TIMP1, soluble CD40L) as well as novel markers such as LP-PLA2 have been developed for development of advanced CVD monitoring panels.
DIAGNOSTICS
SISCAPA technology provides a robust alternative when clinical immunoassays experience interferences (e.g., due to autoantibodies) or are difficult to multiplex.
Serum thyroglobulin (Tg) is used as the primary indicator of thyroid cancer recurrence after thyroidectomy, but clinical immunoassays are subject to interference by endogenous autoantibodies in 20-40% of patients – leading to unacceptable false negative results. Proteolytic digestion of patient samples eliminates these interferences, while SISCAPA enrichment of selected Tg peptides allows highly sensitive measurement by mass spectrometry. SISCAPA Tg assays are provided by numerous clinical reference labs worldwide, and have delivered more than 150,000 accurate patient results to date.
Clinical laboratories routinely measure ~55 different plasma proteins using individual immunoassays. The inherent multiplexability of SISCAPA assays using a common workflow, and the ability to simultaneously measure proteins with abundances differing by more than 1010-fold, permit the creation of panels covering wide areas of clinically-relevant biology. In order to enable use of this established clinical menu in drug trials, we have developed SISCAPA assays for a large fraction of the clinical menu and demonstrated their performance in serum, plasma and whole blood (including dried blood spots).
Other diagnostically useful SISCAPA assays are being developed for specific applications.
PTMs AND VARIANTS
Post-translationally-modifications of proteins influence function (e.g., phosphorylation in signaling systems) and biomarker significance (e.g., glycation of HbA1c).
SISCAPA antibodies capable of binding a specific family of peptides can be developed and used to enrich a target peptide with and without specific chemical modifications (e.g., phosphorylations) or sequence variants (e.g., substitutions distinguishing members of a protein family). For example, we developed a single SISCAPA antibody capable of binding homologous peptides from ERK1 and ERK2, and all versions phosphorylated at either of 2 sites.
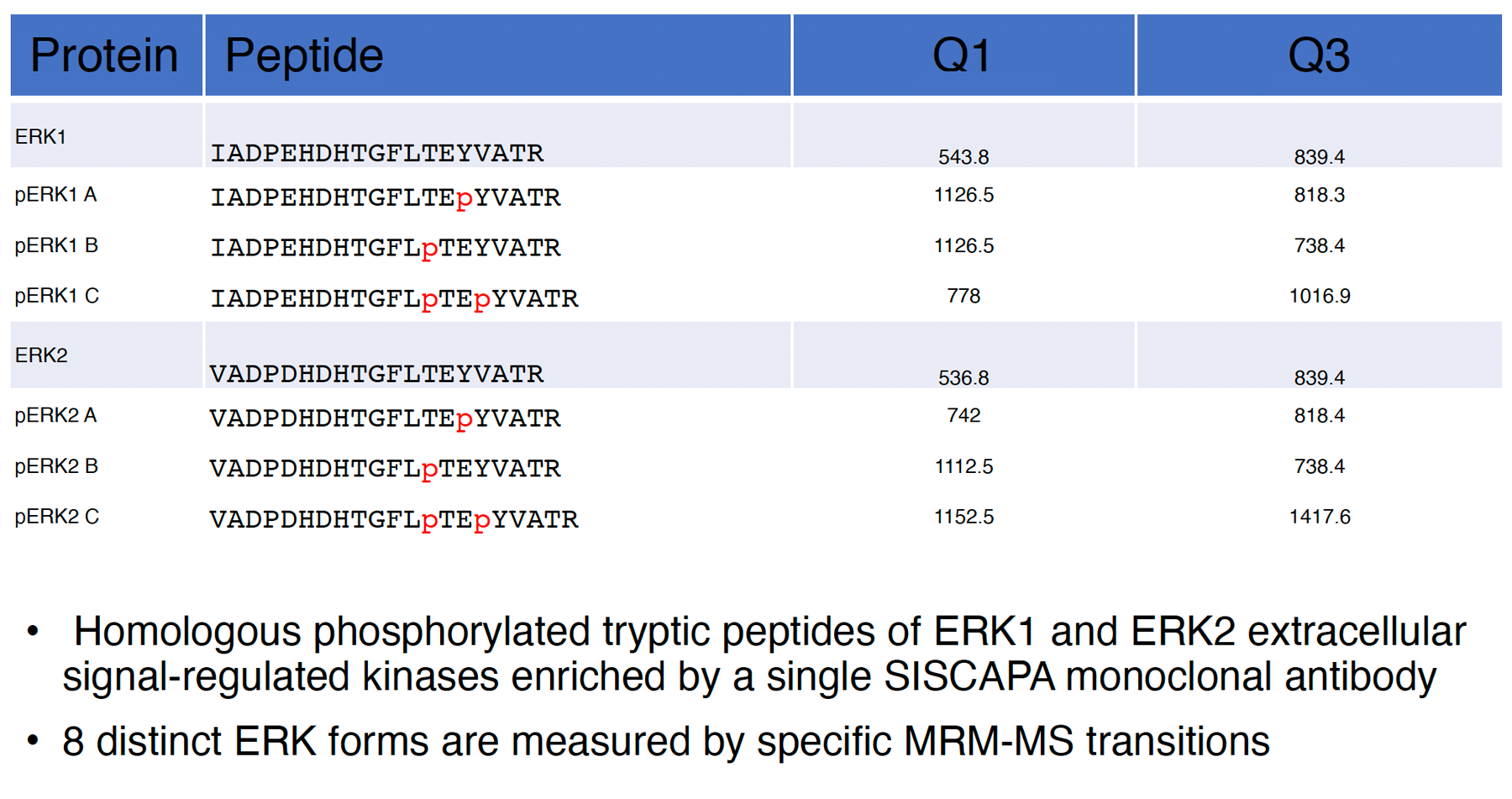
Similarly, SISCAPA assays enable enrichment of peptides with and without adducts formed by reaction with covalent enzyme inhibitors (e.g., reactive drugs), environmental toxicants, and endogenous metabolites (e.g., formation of HbA1c by glycation of HbA, acetylation of tubulin, etc.).
LONGITUDINAL HEALTH MONITORING
Dried blood spot samples
Dried blood spot (DBS) microsamples provide a new window for personalized monitoring of infections, chronic inflammatory disease, and clinical trials of anti-inflammatory drugs.
The specificity and interference-resistance of SISCAPA assays allow their use in the most complex matrices, including whole blood, while the inclusion of proteolytic digestion as the initial step in the workflow makes the process essentially insensitive to denaturation of the sample proteins (e.g., by drying and long-term storage).
We have established the utility of SISCAPA assay panels (e.g., for inflammation, spanning >106-fold in protein abundance) in DBS samples collected over long periods of time, and shown that DBS results are reproducible and closely parallel measurements in venous serum samples. A few proteins show systematic differences between the two sample types, and these are consistent with known biology.
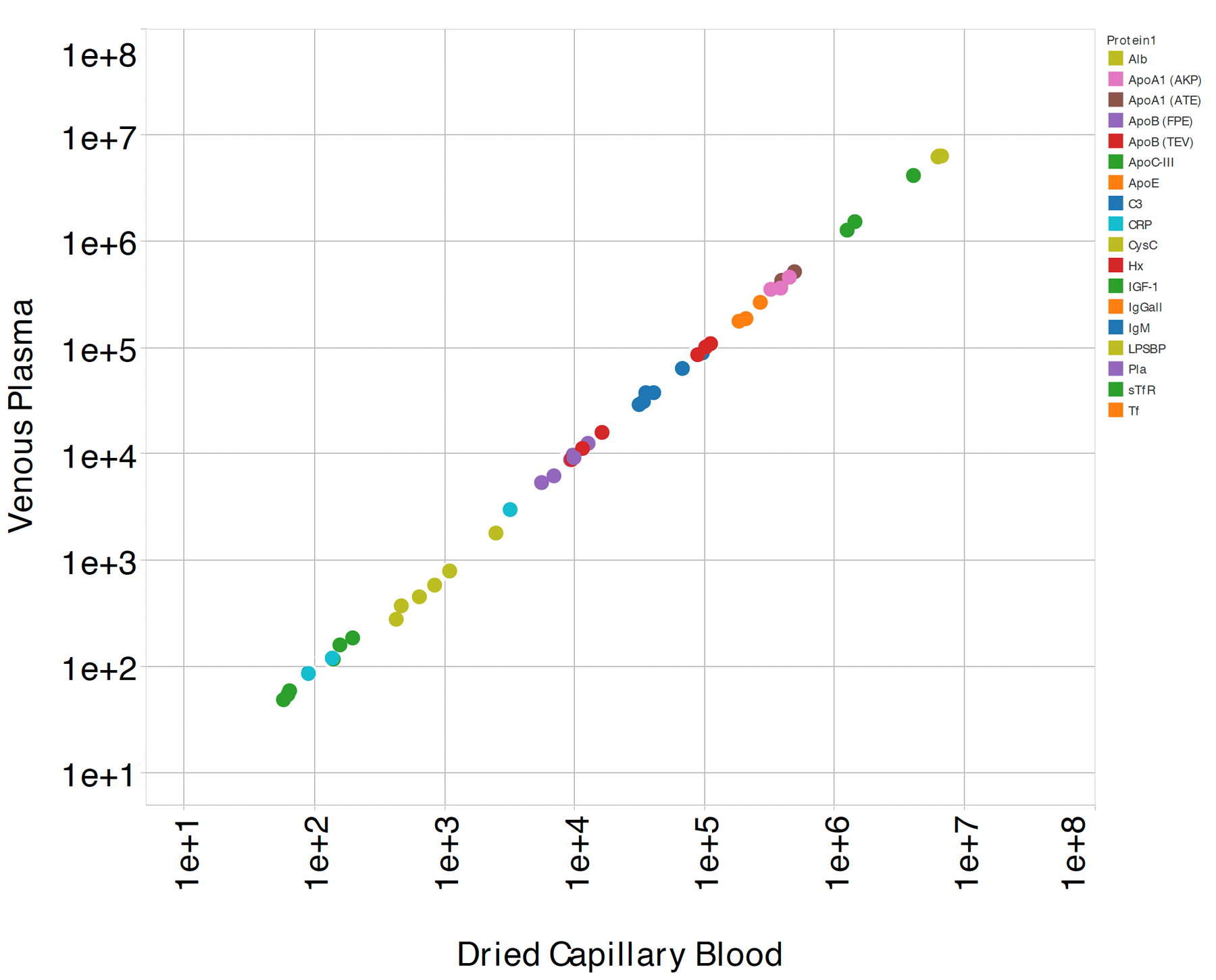
Precision multiparameter tracking of inflammation on timescales of hours to years using serial dried blood spots., L. Anderson, et al, Bioanalysis 12, 937–955 (2020).
Longitudinal health monitoring
DBS sampling enables frequent (e.g., daily) collection, which in turn allows accurate determination of personal baselines for each biomarker and very sensitive detection of deviations from personal baselines. To explore the potential of such high-frequency biomarker tracking and analysis, we have developed a compact, user-friendly sampling system (the SampleDiary™) augmented with an automated contextual information gathering workflow (e.g. data from wearable sensors) described in a separate website (https://www.longitudedx.com/).
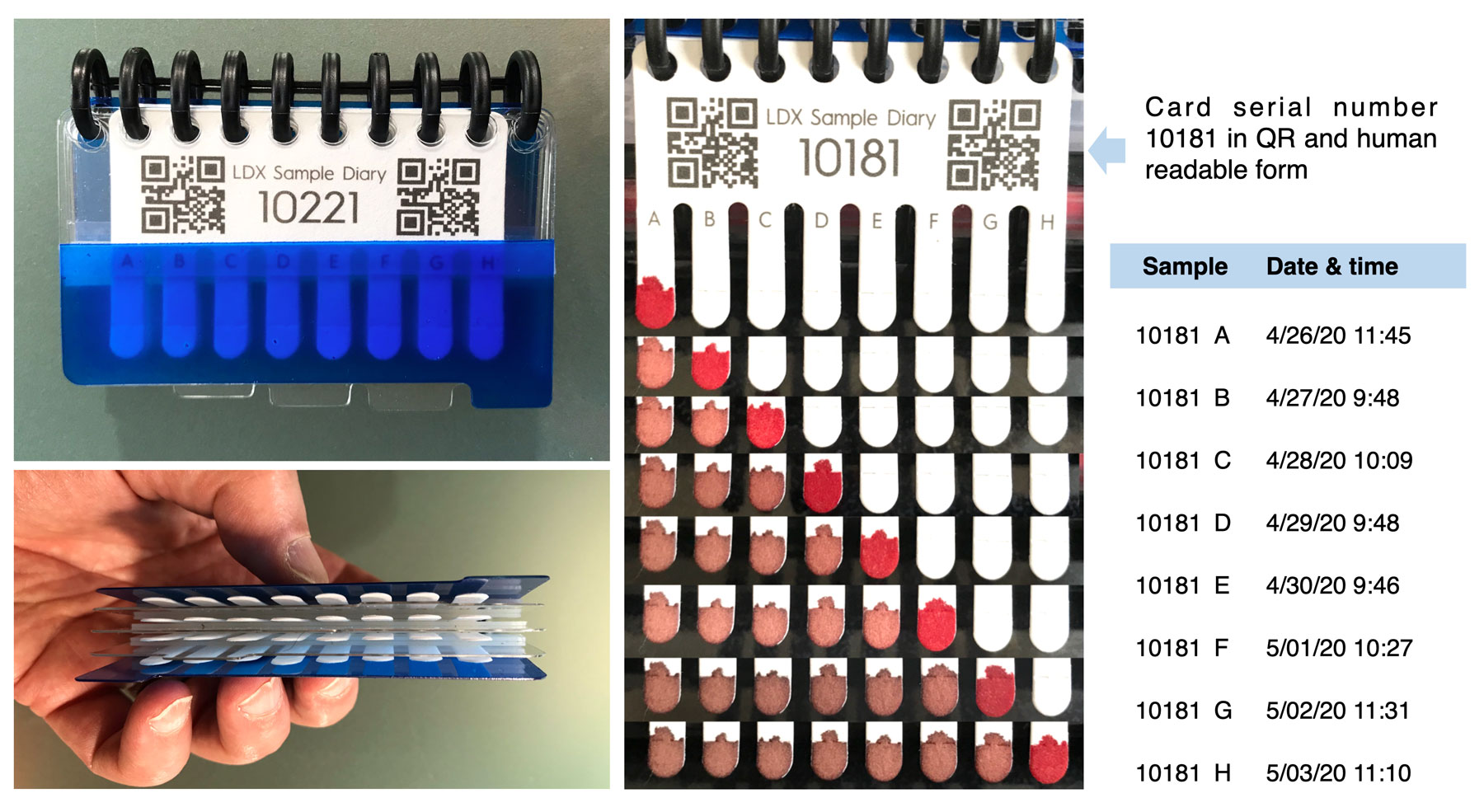 Using DBS microsamples collected at high (e.g., daily) frequency over extended periods, a variety of otherwise-invisible biological responses are observed.
Using DBS microsamples collected at high (e.g., daily) frequency over extended periods, a variety of otherwise-invisible biological responses are observed.